Detailed reviews exist summarizing the different models to use for microfluidic 6 and biological 7 applications, as well as the hardware infrastructure needed to generate high-quality and complex datasets to train ML models. While deeper models can handle more complex datasets and make superhuman inference, they can be data, time, and cost intensive. ML models range in complexity from a simple linear regression to deep neural networks (NNs). Machine learning (ML), the use of trainable statistical models to recognize patterns and predict future behavior, is a promising method to bridge the knowledge gap between experts and end-users and automate the design and operation of microfluidics. 5 While fabrication can be outsourced (albeit at significant cost), microfluidic design and operation can require months to years of multiple “design-build-test” iterations to optimize performance. 4 As recently stated by Battat, Weitz, and Whitesides, wide adoption of microfluidic platforms has been limited by the complexity in the design, fabrication, and operation of custom devices that limit its reproducibility and generalizability. 3 Despite these successes, the impact of microfluidics is mostlylimited single-use cartridges inside of integrated bench-top devices, custom infrastructure within expert labs, or specific applications. Microfluidics has been a core contributor to major technical progress in biotechnology, enabling the development and adoption of commercial platforms such as next-generation sequencing, 1 single-cell RNA sequencing, 2 or droplet digital PCR. 1 Introduction Since its inception, microfluidics has been touted as a revolutionary platform to miniaturize biological and chemical experimentation. Here, we present the current state of machine learning for the design and control of microfluidic devices, its possible applications, and current limitations. Integration of machine learning with microfluidics could significantly expand its adoption and impact. The intuition of microfluidic experts can be captured through machine learning, where complex statistical models are trained for pattern recognition and subsequently used for event prediction. If these barriers were overcome, microfluidics could miniaturize biological and chemical research for non-experts through fully-automated platform development and operation. Despite its ubiquity, the complexity of designing and controlling custom microfluidic devices present major barriers to adoption, requiring intuitive knowledge gained from years of experience. Using this approach, individuals with no training in microfluidics can obtain custom chip designs for their own unique needs in just a few seconds.Microfluidics has developed into a mature field with applications across science and engineering, having particular commercial success in molecular diagnostics, next-generation sequencing, and bench-top analysis. This is one of many different applications for randomly-designed microfluidics in principle, any microfluidic chip that can be simulated could be designed automatically using our method. We also fabricated and tested 16 chips from the library, confirmed that they function as predicted, and used these chips to perform a cell growth rate assay.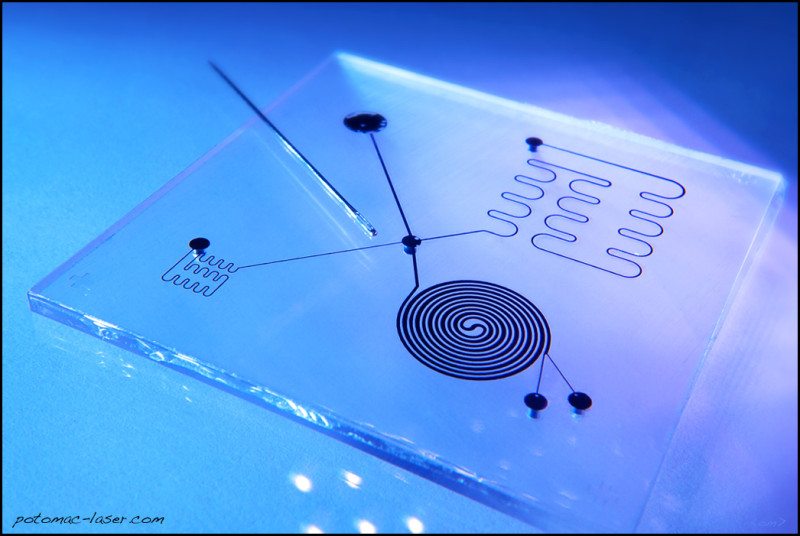 
To demonstrate this functionality, we used our library to select chip designs that generate any three desired concentrations of a solute. The simulation results were then saved to a database which a user can query via to find chip designs suitable for a specific task. We accomplished this by first generating a library of thousands of different random microfluidic chip designs, then simulating the behavior of each design on a computer using automated finite element analysis. In this work we created functional microfluidic chips without actually designing them. 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
